Neurons are highly polarized cells that employ diverse mechanisms to initiate, maintain, and regulate their biological functions, reaching up to 10,000 times more cell surface area than other cell types. In order to establish and maintain distinct neuronal domains (somatodendritic and axonal), specific protein transport mechanisms direct proteins to these compartments so that each domain acquires the appropriate molecular composition.
The endosomal system participates in this complex process by specifically directing membrane components (sorting), internalizing, recycling, or degrading proteins from one domain to another, thereby maintaining the complex neuronal architecture, a defining feature required to sustain physiological processes of neurodevelopment, including migration, polarization, and synaptic function.
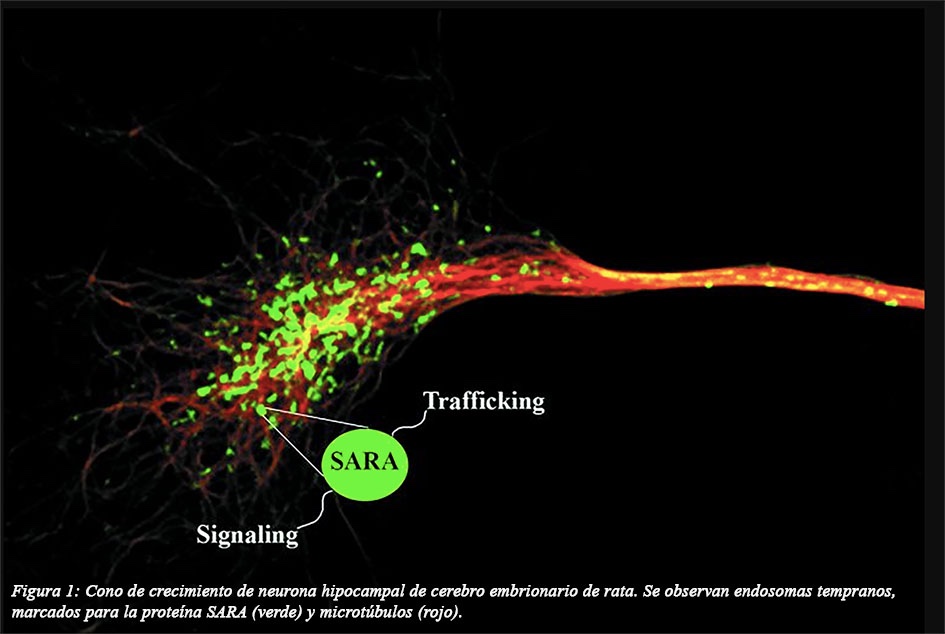
These processes are also tightly regulated by the spatial and temporal expression of specific proteins, as well as by multiple signaling cues, such as the action of TGFβ (Transforming Growth Factor-β), which is required during the development of the central and peripheral nervous systems (CNS and PNS). Smad Anchor for Receptor Activation (SARA) is a signaling protein of the TGFβ pathway that regulates neuronal migration during neocortical development, while its function associated with endosomal trafficking is mediated through binding to early endosomes, participating in the sorting and recycling of membrane proteins (Figure 1).
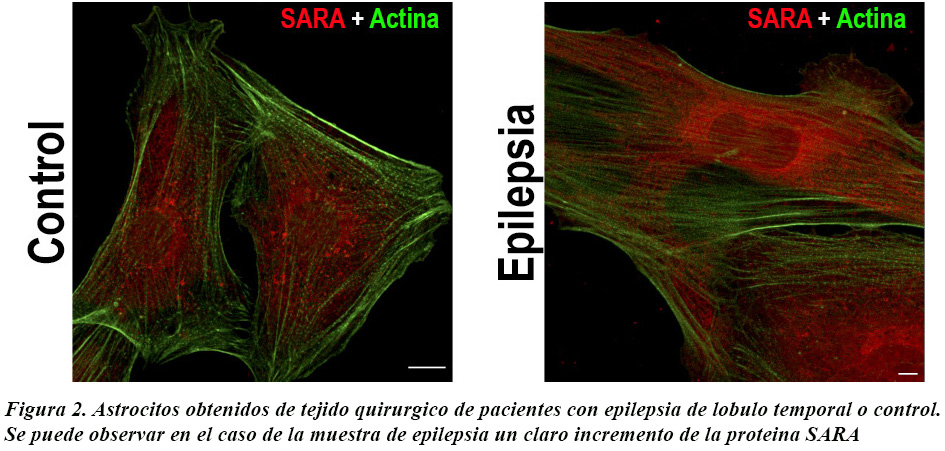
Recently, we have begun to study SARA in astrocytes from patients with temporal lobe epilepsy. Based on our findings, we propose SARA as a potential therapeutic target for patients who are refractory to the classical drugs currently administered to treat this condition (Figure 2).
The projects in our laboratory aim to provide new insights into physiological and pathological processes of neurodevelopment, specifically by identifying and/or characterizing molecules and mechanisms that: (1) participate in the endosomal system and regulate trafficking in neurons, as well as their role in the morphological changes observed during neurite growth; (2) regulate TGFβ signaling, enabling successful axonal establishment and growth; and (3) contribute to pathological processes, specifically temporal lobe epilepsy.
In our experiments, we employ a range of cellular and molecular biology techniques, high-resolution microscopy, and biochemical approaches. We work with murine models through neuronal cultures (in vitro), as well as by manipulating gene expression during mouse brain development using the in utero electroporation technique (in vivo). We also use experimental models of status epilepticus and cultures of human astrocytes derived from patients with epilepsy.
COMPETITIVE RESEARCH FUNDS
- 2021-2023 Transcription factors of CREB3 family: modulation of secretory pathway and implications in development and regeneration of the nervous system. National Research Council, CONICET. PI: Cecilia I. Alvarez; Co-PI: Cecilia B. Conde
- 2021-2024. Functional analysis of SARA in newest endosomal vesicles in nervous system: potential implications on protein sorting during neurodevelopment. (PICT 2020-01268). National Agency of Science and Technology Promotion (ANPCyT-FONCyT). Argentina. PI
- 2023-2027. Functional analysis of the SARA protein in smooth muscle cells and in a model of vascular remodeling: physiological and pathological implications. (PICT 2021-00280). National Agency of Science and Technology Promotion (ANPCyT-FONCyT). Argentina. PI: Cecilia B. Conde; Co-PI: Melina Musri.
- 2023 IBRO Collaborative Research Grants. SARA and Smurf2/Smad7 complex activity in temporal lobe epilepsy: implications on TGF pathway modulation. Dr. Conde C (INIMEC-CONICET-UNC) and Dr. Kawauchi T (Kyoto University, Japan)
PUBLICATIONS (LATEST FIVE YEARS)
- Natali L, de la Cruz-Thea B, Godino A, Conde C, Peinado VI, Musri MM. A Comprehensive Review of Epigenetic Regulation of Vascular Smooth Muscle Cells During Development and Disease. Biomolecules. 2026 Jan 21;16(1):173. doi: 10.3390/biom16010173. PMID: 41594714.
- Conde C, Bates EA, Garcia C, Lazo OM. Editorial: Neuronal cytoskeleton and GTPases in health and diseases. Front Cell Dev Biol. 2022 Sep 15;10:1025527. doi: 10.3389/fcell.2022.1025527. PMID: 36187477.
- Capelluto DGS, Conde CB, Tumbarello DA, van den Bogaart G. Editorial: Signaling Proteins for Endosomal and Lysosomal Function. Front Cell Dev Biol. 2021 Dec 16;9:821719. doi: 10.3389/fcell.2021.821719. PMID: 34977050.
- Urrutia PJ, Bodaleo F, Bórquez DA, Homma Y, Rozes-Salvador V, Villablanca C, Conde C, Fukuda M, González-Billault C. Tuba Activates Cdc42 during Neuronal Polarization Downstream of the Small GTPase Rab8a. J Neurosci. 2021 Feb 24;41(8):1636-1649. doi: 10.1523/JNEUROSCI.0633- 20.2020. Epub 2021 Jan 21. PMID: 33478991.
- Rozés-Salvador V, González-Billault C, Conde C. The Recycling Endosome in Nerve Cell Development: One Rab to Rule Them All? Front Cell Dev Biol. 2020 Dec 10; 8:603794. doi: 10.3389/fcell.2020.603794. PMID: 33425908.
- Rozés-Salvador V, Wilson C, Olmos C, Gonzalez-Billault C, Conde C. Fine-Tuning the TGFβ Signaling Pathway by SARA During Neuronal Development. Front Cell Dev Biol. 2020 Sep 3; 8:550267. doi: 10.3389/fcell.2020.550267. PMID: 33015054.
- Siri SO, Rozés-Salvador V, de la Villarmois EA, Ghersi MS, Quassollo G, Pérez MF, Conde C. Decrease of Rab11 prevents the correct dendritic arborization, synaptic plasticity and spatial memory formation. Biochim Biophys Acta Mol Cell Res. 2020 Sep;1867(9):118735. doi: 10.1016/j.bbamcr.2020.118735. Epub 2020 May 7. PMID: 32389643.
- Complete and updated list available here.
TEAM